Research
Research Projects
On a path to precision child health cardiovascular medicine, the Neonatal/Congenital Heart Program consists of three major complementary research projects:
I. Congenital Heart Defects Genetics & Genomics
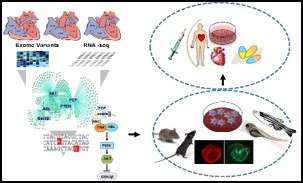
Developmental and structural abnormality of the heart affect approximately 1% of live births and is a major source of childhood morbidity and mortality. The underlying mechanism is known to be multifactorial, involving genetic, epigenetic and/or environmental contributors. However, the majority of CHDs are yet to be clearly diagnosed for the underlying causing factors, and even less with effective therapies. Therefore, a clear dissection of the underlying mechanism is the most powerful step to establish individualized preventive strategies and patient tailored therapies. To that end we Implemented a systems based approach to investigate congenital heart defect mechanism as a continuum from phenotypic characterization in the clinic, to molecular genetic dissection at the bench. Our team integrates the power of next generation sequencing platforms, including family-based Exome sequencing and RNA-seq data, to identify novel candidate genetic variants, environmental causes and to test their potential interactions in zebrafish and patient specific cell line. Ultimately, our goal is to identify new diagnostic and preventive approaches for congenital heart defects.
II. Chamber Specific Transcriptome Programming in Newborn Hearts with Congenital Heart Defects
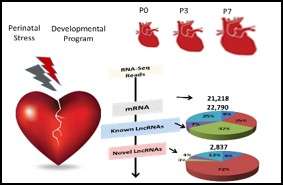
Our current knowledge on cardiac development is largely built upon insights to transcriptional network driving cardiomyocyte lineage determination in fetal heart. However, the molecular basis for chamber specific programming during perinatal maturation is vastly understudied. Despite of the significance of external stress factors, such as hypoxia on congenital heart defects pathogenesis, the current focuses remain at gene factors rather than the noncoding regulatory elements and stress signaling. In this project, we employed unbiased transcriptome analysis in neonatal mouse hearts and conducted extensive expression and functional analysis leading to discovering hundreds of novel lncRNAs that are conserved in human infantile heart, serving as putative molecular biomarkers for congenital heart defect phenotypes. We also identified Wnt11 as a key molecule implicated in chamber specific myocyte proliferation regulation under hypoxia stress and contributes to hypoxia-induced developmental changes in cyanotic congenital heart defects. Our team employs a novel mouse model and molecular biology tools in cell culture system to investigate the signaling mechanism of Wnt11 regulation of chamber specific development in response to hypoxia stress. The long-term goal of our efforts is to identify chamber specific therapies for the infants with congenital heart defects.
III. Congenital Heart Defect-BioSpecimen Core Resource (the CHD-BioCore) at UCLA
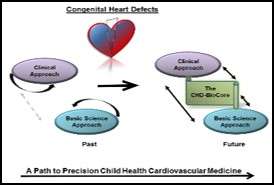
Access to high-quality biological specimens from large and well-defined patient cohorts is a rate-limiting step in any effort to study the patho-mechanisms of congenital heart disease. Therefore, we established the UCLA-CHD-BioCore. The CHD-BioCore consists of three central and complementary components:
- CHD-Clinical Data Registry
- CHD-Bio Banking Repository
- CHD-Genomic Data Depository
We are currently enriching CHD-BioCore with patient specific-induced Pluripotent Stem Cell lines (iPSCs) in collaboration with Dr. Joseph Wu, Director of the Stanford Cardiovascular Institute. Subsequently, the cells will be used to derive a patient specific cardiomyocyte or other cell types that will serve as a powerful system to examine disease mechanisms and patient tailed therapies. By synergizing expertise from a collaborative team with combined clinical and research expertise, the BioCore will greatly facilitate scientific discovery and clinical translation and serve to promote collaboration with CHD consortium network member institutes nationwide.
Fellowship Research
One position is currently open for fellowship research