Dr. Corey Arnold Lab

The UCLA Computational Diagnostics Lab (CDx) uses machine learning to better understand disease through the discovery of predictive computational phenotypes from the fusion of information across diagnostic disciplines. We seek to translate our results to practice through the construction of novel clinical tools and workflows. CDx is joint effort between the Department of Radiological Sciences and the Department of Pathology & Laboratory Medicine, and is affiliated with the Medical Imaging & Informatics (MII) group and the Center for Telepathology & Digital Pathology.
Projects
Our projects span several diseases and modalities, but share the common theme of incorporating machine learning. Much of our work focuses on the analysis of radiology and pathology image using deep learning techniques. We currently have NIH-funded projects in prostate cancer, acute stroke, and mHealth.
Predicting the Presence of Clinically Significant Prostate Cancer using Multiparametric MRI and MR-US Fusion Biopsy
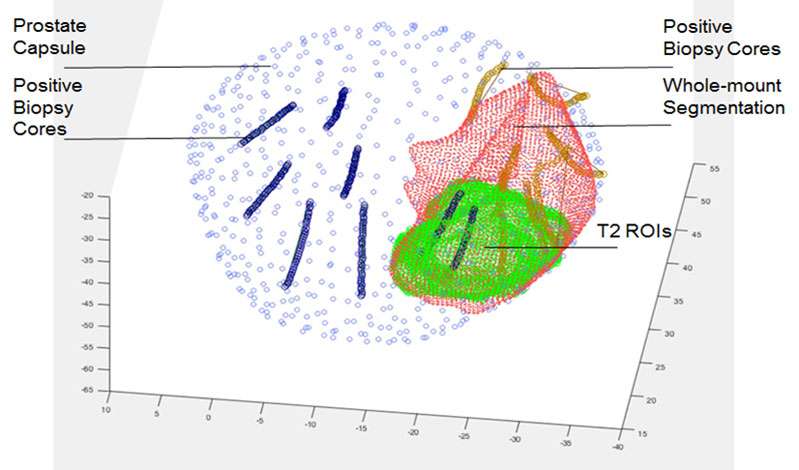
Prostate cancer is the second leading cause of cancer death in American men, accounting for 26% of new cancer diagnoses and 9% of cancer deaths in men. Active surveillance, radical prostatectomy and radiotherapy are commonly used treatments for clinically localized prostate cancer. However, current risk stratification methods cannot be used effectively to avoid subjecting patients with clinically indolent cancers to unnecessary interventions, causing significant morbidity and cost.
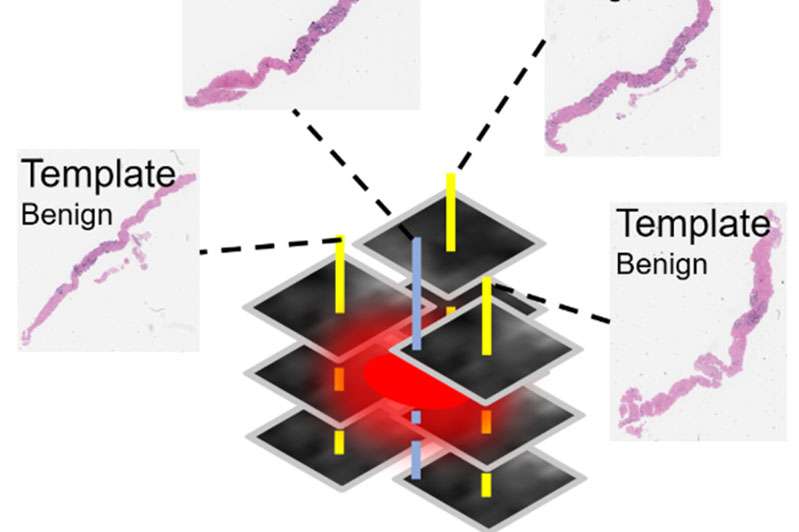
The research objective of our R21 is to develop novel techniques using multiparametric magnetic resonance imaging (mp-MRI) and MRI-ultrasound (US) fusion guided biopsy data that provide discriminatory power in distinguishing indolent versus clinically significant prostatic adenocarcinoma based on non-invasive imaging. We are implementing a multi-instance learning (MIL) based convolutional neural network (CNN) model for clinical prostate mp-MRI sequences to generate new quantitative imaging features representative of the underlying tissue. Our MIL-CNN model accommodates ground truth labels from pathology whole mount specimens, as well as MRI-US fusion biopsy results. Hierarchical CNN features will be used to predict voxel-level cancer suspicion, thereby enabling a novel method for performing “imaging biopsy.” Finally, voxel-level suspicion maps will be aggregated into patient-level quantitative imaging biomarkers and combined with clinical data to create a multimodal nomogram for performing risk stratification.
NIH National Cancer Institute, R21 CA220352
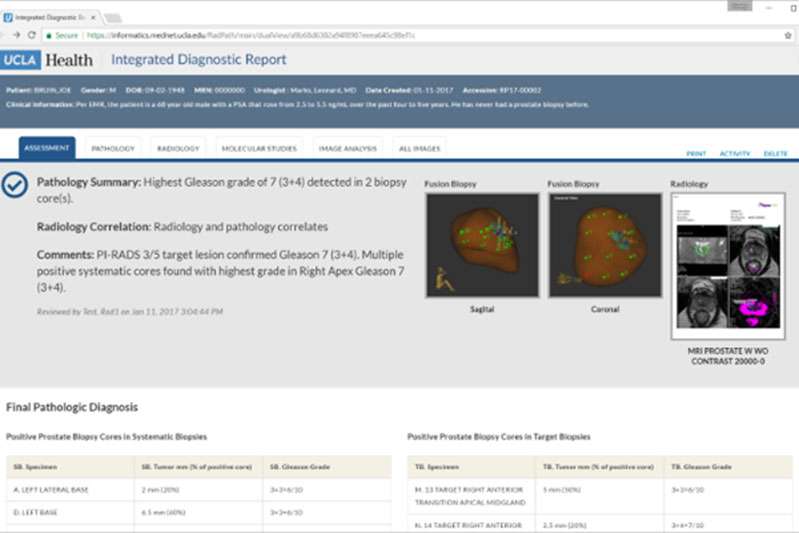
Integrated Diagnostic Reporting
We have developed a reporting platform for improving diagnostic accuracy and efficiency through the integration of clinical data with AI support. The system brings together clinical, imaging, pathology, and molecular data into a single view, allowing physicians to more efficiently review a patient’s case with a comprehensive information set. The platform is currently in use at UCLA for cancer diagnosis with over 10,000 cases completed.

Resources
Our lab maintains several project resources. For compute, we have an NVIDIA DGX-1 GPU processing server that has high-speed connections to our data server. We also maintain a Leica/Aperio GT450 whole slide scanner for digitizing histology slides.